
Contact information
peijun.zhang@strubi.ox.ac.uk
peijun.zhang@diamond.ac.uk
Professor Peijun Zhang Research
Henry Wellcome Building of Genomic Medicine
Colleges
 |
| Complete all-atom model of HIV-1 capsid (Nature 497:643, 2013, featured on Nature Cover). |
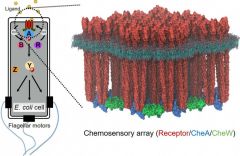 |
| Bacterial chemosensory arrays (Elife 4:e08419, 2015) |
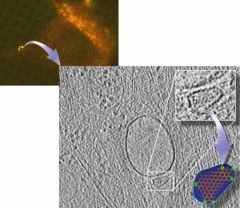 |
| Direct visualization of HIV-1 infection using correlative microscopy (Structure 19:1573, 2011) |
Peijun Zhang
Professor of Structural Biology and WELLCOME TRUST INVESTIGATOR
Structural Biology of Human Pathogens
Our research is aimed at an integrated, atomistic understanding of molecular mechanisms of virus and bacteria infections by developing and combining novel technologies for high-resolution cryoEM and cryo-electron tomography with complementary computational and biophysical/biochemical methods. Current research efforts in our lab are directed to:
HIV-1 capsid assembly, maturation, and interactions with host cell factors
Retroviruses, such as human immunodeficiency virus 1 (HIV-1), contain mature conical capsids that enclose the viral RNA genome, enzymes and accessory proteins. The assembly and stability of the viral capsid are critical to the viral replication life cycle. Structures of the building blocks of the capsid assembly were determined to atomic level, the mechanisms of capsid assembly and disassembly during a productive infection, however, remain unclear. Such information is essential for the development of therapeutic drugs that target viral capsid. More importantly, the surface of HIV-1 capsid serves a primary interaction interface between the virus and the host cell. Many host defense proteins have been identified to interact with the viral capsid and block HIV-1 infection. Yet, very little is known about their precise recognition and interactions, and thus mechanisms of inhibition. We are developing cutting-edge cryoEM technologies that bring unprecedented resolution and enable in situ structures of HIV-1 and in complex with host proteins, such as CypA, TRIM5α, TRIMCyp, CPSF6 and MxB, to decipher their underlining functional roles
Bacterial chemotaxis sensory arrays
Bacteria use chemotaxis signaling pathways to monitor their environment and respond appropriately to change, which is crucial for colonization and infection for bacterial pathogens. The essential core signaling unit comprises transmembrane receptors, a histidine kinase CheA, and a coupling protein CheW. Remarkably, bacteria accomplish the extraordinary gain and cooperativity in chemotaxis signaling by arranging a few hundred core signaling units into higher order arrays localized at the cell pole. We aim to determine the precise molecular mechanisms of chemotaxis cooperative signaling using high-resolution cryoEM and cryoET in combination with site-directed mutagenesis and computational modeling. Our long-term goal is to develop plausible molecular models, at atomic resolution, for the entire signaling pathway by assembling structural “snapshots” of the signaling states.
CryoEM technology development
Driven by biological questions and inspired by the bottlenecks we have to overcome, we devote significant efforts to the advancement of novel cryoEM methods and technologies. We are working on a wide spectrum of technical advances that will be essential to realize the promise of imaging cells and tissues at molecular resolutions, including correlative microscopy, cryo-FIB/SEM and high resolution sub-tomogram classification and averaging.
Major funding
Wellcome Trust Investigator Award: Molecular mechanisms of HIV-1 restriction by capsid-sensing host cell proteins
BBSRC project grant: Assembly and Dynamics of Bacterial Chemosensory Signaling Arrays
NIH/NIAID: University of Pittsburgh Center for HIV Protein Interactions (PCHPI)-CryoEM Core
ERC AdG grant: Molecular choreography of bacterial chemotaxis signalling
Recent publications
-
Editorial overview: Cryogenic electron microscopy (cryoEM).
Journal article
Rohou A. and Zhang P., (2024), Current opinion in structural biology, 84
-
ChAdOx1 COVID vaccines express RBD open prefusion SARS-CoV-2 spikes on the cell surface.
Journal article
Ni T. et al, (2023), iScience, 26
-
Structure of native chromatin fibres revealed by Cryo-ET in situ.
Journal article
Hou Z. et al, (2023), Nature communications, 14
-
Intrinsically disordered CsoS2 acts as a general molecular thread for α-carboxysome shell assembly.
Journal article
Ni T. et al, (2023), Nature communications, 14
-
Multidisciplinary studies with mutated HIV-1 capsid proteins reveal structural mechanisms of lattice stabilization.
Journal article
Gres AT. et al, (2023), Nature communications, 14
-
Structure of the native chemotaxis core signaling unit from phage E-protein lysed E. coli cells.
Journal article
Cassidy CK. et al, (2023), mBio
-
Molecular architecture and conservation of an immature human endogenous retrovirus.
Journal article
Krebs A-S. et al, (2023), Nat Commun, 14
-
Cryo-electron tomography to study viral infection.
Journal article
Graham M. and Zhang P., (2023), Biochem Soc Trans
-
Neutral sphingomyelinase 2 is required for HIV-1 maturation.
Journal article
Waheed AA. et al, (2023), Proc Natl Acad Sci U S A, 120
-
Molecular architecture and conservation of an immature human endogenous retrovirus.
Journal article
Krebs A-S. et al, (2023), bioRxiv
-
Cryogenic electron ptychographic single particle analysis with wide bandwidth information transfer
Journal article
Pei X. et al, (2023), Nature Communications, 14
-
ChAdOx1 COVID vaccines express RBD open prefusion SARS-CoV-2 spikes on the cell surface
Preprint
Zhang P. et al, (2023)
-
Recent structural advances in bacterial chemotaxis signalling
Journal article
Riechmann C. and Zhang P., (2023), Current Opinion in Structural Biology, 79
-
Structure of the Native Chemotaxis Core Signalling Unit from E-gene lysedE. colicells
Preprint
Cassidy CK. et al, (2023)
-
Mechanisms of E. coli chemotaxis signaling pathways visualized using cryoET and computational approaches.
Journal article
Hadjidemetriou K. et al, (2022), Biochemical Society transactions
-
Cryo‐EM
structures of perforin‐2 in isolation and assembled on a membrane suggest a mechanism for pore formation
Journal article
Yu X. et al, (2022), The EMBO Journal
-
Structure and activity of particulate methane monooxygenase arrays in methanotrophs.
Journal article
Zhu Y. et al, (2022), Nature communications, 13
-
Application of super-resolution and correlative double sampling in cryo-electron microscopy.
Journal article
Sheng Y. et al, (2022), Faraday discussions
-
Structure and assembly of cargo Rubisco in two native α-carboxysomes.
Journal article
Ni T. et al, (2022), Nature communications, 13
-
Cryo-electron Tomography Remote Data Collection and Subtomogram Averaging.
Journal article
Sheng Y. et al, (2022), Journal of visualized experiments : JoVE


