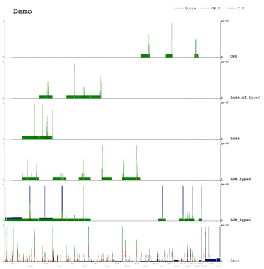
Try the demo files or have a look at some published work using this site to get a better idea:
Actions
- Plot Data
- Graphically depicts complete data for selected chromosomes. Can be filtered by Chromosome, Type of Events, cytoband name and overlap.
- Table File
-
| Beware this can take few minutes with larger array. |
| Note: The file should be TAB delimited with following header and unlimited number of rows: |
| PatientID |
Chr |
Type |
Start |
End |
Comment |
|
A few sample datasets are available:
- Mini dataset: Contains a few lines of demonstration as a text file demo_mini.txt
- Realistic dataset: Contains a longer demonstration as an Excel (2004) file demo.xls.
Note: This file will take about a minute to process
- Gene File
-
The gene file contains information about genes to be plotted in the chromosome view.
It shall contain information about the gene location and name with a header containing the following fields:
- Name The label to be used for this gene
- Chromosome The chromosome where the gene is located (chr1, chr2, ...)
- Start Location The Location in bp
- End Location The End location in bp. This is optional as if missing a segment (eventually more visible) will be drawn
Note that the file has to be in text format
- Configuration File
-
The Configuration file contains some parameters to run the program:
- Graphics: Various graphical parameters
- Offset: The localisation of each chromosome for the summary plot
- colors: The colors used for each type of event
- stain: The color scheme for the cytobands
- summary_bbx, summary_bby: Size of the bounding box defining the summary page
- event_size, event_skip: width and skip of events in the sumary view
- block_lim, block_size: Limit before collapse of identical events, and the corresponding size
- Misc: Other parameters of importance
- Conv: The conversion command from Encapsulated Postscript (EPS) to the other formats
More information about the conversion options is available from ImageMagick
Sections are defined between square brackets, '[]', and comments start with '#' or ','
A demonstration configuration file is available
Filters
Select Chromosomes:
Select a specific chromosome number or all the chromosomes at once by selecting "Genome-Wide Summary"
Select visible parameters:
Information on Plot
Select whether you want to see some particular information on the plot
- Overlap of Events:
- Show the overlap of a specific Event type
- Collapse Events in summary:
- Allow identical events across samples to be merged in summary.
Details can be modiofied with the configuration file.
- Cytoband Name:
- Show the name of the cytoband inside the chromosome if possible
- Show by individuals:
- The Events are presented by a fixed set of individuals, with overlapping events.
Currently only implemented for the genome-wide view
- Highlight LOH:
- If the string "LOH" is found in the event name, then this section will contain extra highlighting.
Suggest to set the same color for the event with and without LOH with the Configuration file.
- DGV:
- Known CNV from the Database of Genomic Variants can by included in the plot.
Because of the diversity of source for this resource it is possible to add a filter for a given pulication and/or Author:
The filter works as an AND operator of ":" separated fields. For example Affymetrix:Conrad will provide all variants found with Affymetrix platform AND found in Conrad et al.
- Plot Profile:
- Show the selected score on the chromsome and new summary view
- Show Score:
- Show the Score in the above profile
- Show GW Poisson-Bionomial:
- Show the Genome-Wide Poisson-Bionomial in the above profile
- Show Chr. Poisson-Bionomial:
- Show the Chromosome-Wide Poisson-Bionomial in the above profile
Build
Three builds are available to match the events to the cytobands: Build35 (hg17), Build36 (hg18) and Build37 (hg19).
DGV references will be set automatically according to these three builds.
Build 37 is the default.
Output Image Format
Five possible output formats are available for the plots:
By default Encapsulated PostScript (EPS) and are provided JPG (images shown).
But it is possible to generate TIFF, PDF and PNG format as well.
Beware that the more format generated the longer it will take to process.
Output
Multiple output are possible depending on the chosen parameters:
Summary
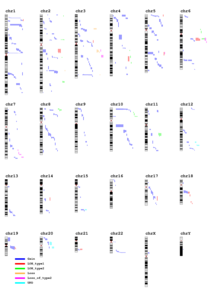
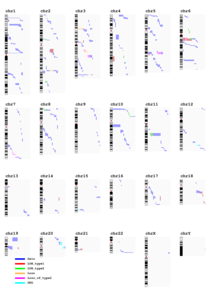
- A Summary genome-wide plot with all events and cytobands are presented by individual or per event.
It does not contain information about overlap, gene or DGV.
It is available in as many format as selected
- A single PDF file is provided with all generated figures
- A single Excel-type file is provided with all generated overlapping table
Per chromosome
For each chromosome there are essentially two type of output:
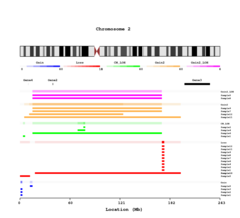
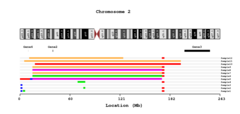
- Plot
From bottom to top, the events are plotted with specific color per event or per individual, with the name of the sample on the side.
If the Highlight option is selected a dark line is showing region with LOH.
If selected the overlap of each event type is given with darker color corresponding to more common regions.
If selected the filtered DGV regions are shown in dark yellow.
If given, the gene list, with their corresponding name are drawn.
If the overlap is selected, the legend with color and intensity information for each even is shown.
The cytobands for the chosen chromosome.
Other format for the plots are available according to the selection.
- Overlap table
If the overlap calculations and the "Show overlap details" were selected a table will be provided in both Excel friendly format on directly on the site with a summary of the highest count.
The columns are as follow:
- Chromosome: chromosome presented
- Type: Type of event concerned as provided in Input file, usually Gain, Loss
- Start: Start position of the segment
- End: End position of the segment
- Count: Number of samples carrying this given type of event at this location
- Score: Proportion of samples carrying this given type of event at this location (0..1)
- GW_P: Poisson-binomial test for overlap of the event across the individuals at this location, given the proportion of the genome affected.
- C_P: Poisson-binomial test for overlap of the event across the individuals at this location, given the proportion of the chromosome affected.
- UCSC: Link of the region to the UCSC genome-browser to have the overlap in the context of further information
log files
Both Standard Output and Standard Error from the Python command are available for inspection.
These should help investigating potential reason for failure to process as expected.
4. Command line version
This page propose a convenient graphical interface to a Python based software.
However, the Python engine is available directly to allow batch runs as well as very specific tailoring.
To install GREVE on your own machine you will need the following pieces:
- Python Packages:
- xlrd
- Library for developers to extract data from Microsoft Excel (tm) spreadsheet files.
- vplot
- A vector plotting program written in object-oriented Python.
- rpy2
- A low-level interface to R, a proposed high-level interface, including R-like structures and functions.
- DGV variants and indel files matching your preferred build from the repository:
- R Package:
- poibin
- This package implements both the exact and approximation methods for computing the cdf of the Poisson binomial distribution.
- Image manipulation software:
- ImageMagick
- ImageMagick is a software suite to create, edit, compose, or convert bitmap images. It can read and write images in a variety of formats (over 100).
Use ImageMagick to resize, flip, mirror, rotate, distort, shear and transform images, adjust image colors, apply various special effects, or draw text, lines, polygons, ellipses and Bezier curves.
- GREVE:
- Retrieve the GREVE tarball file.
- Decompress the file with
This should create a folder with both necessary and optional example files
- Run a command with the demo files such as:
- GREVE.py -h
- GREVE.py --jpg -c chr2 -c chr4 --ind -s --loh demo_mini.txt
- GREVE.py -w -n -f default.cfg -g gene.list demo.xls
Do not hesitate to contact the author to share some extension of yours.
5. F.A.Q.
- Nothing is showing. What is going on?
- You need to make sure that you follow the 2 possible formats described above: TAB separated text file, Excel file.
Furthermore make sure you filename contains only 1 ".".
If you use Excel format, make sure it is 97-2004 format, rather than the latest.
Finally make sure that you have setup enough colors in the configuration file for the number of event type you are using.
- Processing is slow. Is it normal?
- Depending on the size of your input file, and the action required, the process can take few minutes.
You might want to skip the costly verlap definition, reduce the number of file type generated, or the number of chromosomes analysed.
- I do not like to color choice. Can I modify it ?
- Yes. The choice of color is made in the order of the type found in your list.
You can modify it by giving your own configuration file, selection your own color in Red-Green-Blue (RGB) code.
- My bars go over to the next chromosome. Can I change this behaviour?
- Yes. First I would recommend to tick the "Collapse Event in summary" box to put together recurrent events.
Furthermore, the respective position of the chromosome, the size of the bari, the distance between bars are all configurable.
You can modify it by giving your own configuration file.
- There are a lot of overlap on the centromeres and telemores. Is this real?
- Unlikely. There is little information in terms of markers in those regions and their recurrent calling is probably an artefact of the data and calling method.
It is recommended to filter the list of CNV before submitting them to GREVE.
- Can I avoid sorting my event and see the Events per individual?
- Yes. There is an option to plot the Events per individual rather than per event (--ind). In this case the order of the input file is kept across all chromosomes.
- Why is the indivual plot not showing on the chromosome level?
- Currently only the Genome-View has got the individual plot implemented. The chromosome is in the ToDos list.
- My gene list does not seem to be loaded or shown. Is this feature not working?
- The gene list is limited to about 50,000 features because of PHP restictions.
This should be sufficient for gene illustration, especially with the names showing. It is however possible to include more features by using the command line directly.
- I cannot see the LOH events despite checking the box. What is happening?
- LOH is represented by a thin line around the normal box. Because of the various formats it can be difficult to see on the screen (jpeg).
However it should be clearly visible in the EPS and PNG formats.
- What is the Poisson Binomial test?
- The Poisson Binomial uses the proportion of each individual carrying a given event, genome-wide and chromosome-wide, to test how likely it is they overlap N times.
If the probabilities for each individual were equal, it would correspond to the usual Binomial distribution.
- Which score should I use?
- There are 4 scores reported for each segment of each type. They can be classified into 2 subgroups:
- Test of existence against the hypothesis of no event in the control group. Because of the NULL hypothesis, there is no point in reporting a p-value for this test.
- Count: The exact number of individual carrying the given type at the location
- Score [0..1]: The Proportion of individual carrying the given type at the location. This is simply the previous count divided by the total number of individuals to ease comparison between studies.
- The Poisson-Binomial is based on the hypothesis that many events reflect noise. Therefore this test the probability that a given type of event overlaps that many times across individuals.
- GW_P: The probability used for each individual is the proportion of this sample to carry this type of event, i.e. the sum of all such events across the genome.
- C_P: The probability used for each individual is the proportion of this sample to carry this type of event, i.e. the sum of all such events across the examined chromosome.
- Is GREVE used at all beyond the author?
- Yes, the Geomap shows the usage around the world, turning into many known publications
- The GUI is great, but I'd like to run it myself. Is there a command line version?
- Yes this is essentially a web interface to a python based software. You can find the details above
- Whaoo this tool is fantastic !!! Is it published?
- Yes. It has been accepted for publication by Bioinformatics
- My question is not part of the FAQ. How shall I do?
- Please contact the author who will be delighted to attempt solving your problem
6. Recent Changes.
- March 2013
- Create a single PDF file with all images across the genome and chromosomes,
- Create a single Excel file of overlap across the genome rather than one per chromosome,
- Display the scores in a single plot across the genome, in a style similar to a Manhattan plot with segments.
- Include the Manhattan style plots in the regular chromosome view,
- Minor bug correction for non standard Build 37 plots and invalid system error.
- September 2012
- GREVE is accepted for publication in Bioinformatics
- Allow for the Excel format not to have a Commentary column
- July 2012
- Port GREVE's web interface to a newi, faster, server, with more recent PHP version.
- Modify the table output to allow some statistical measure over overlap probability.
- Allow for inclusion of DGV data.
- Add Statistical measure for Proportion and Poisson-Binomial.
- Add Location of users into a map.
- Modify the F.A.Q. accordingly.
- May 2012
- Create this entry
- Allow for 'Space' in the input, Configuration and Gene filename.
- Solve limitation of number of event by increasing the number of colors available
- Fix TIF format issue incompatible with Windows by using ImageMagick "-type truecolor" option.
- Identify unspecified Excel issue to be due to incompatibility with "recent" version of Excel. Need to save as "Excel 97-2004 format".
- Modify the F.A.Q. accordingly
7. Still To Do.
- Add pair testing to the web-version rather than just hard-coded.
- Create a version for non-human CNV (in process through collaboration)
- Allow for newer Excel format by replacing the xlrd package
- Make Circos plot for the summary.
- Allow parameters on the Configuration filei, such as bar thickness, to be modified on the main page.
Last updated on November 2012
Please contact Jean-Baptiste Cazier for details
 Genomic Recurrent Event ViEwer
Genomic Recurrent Event ViEwer